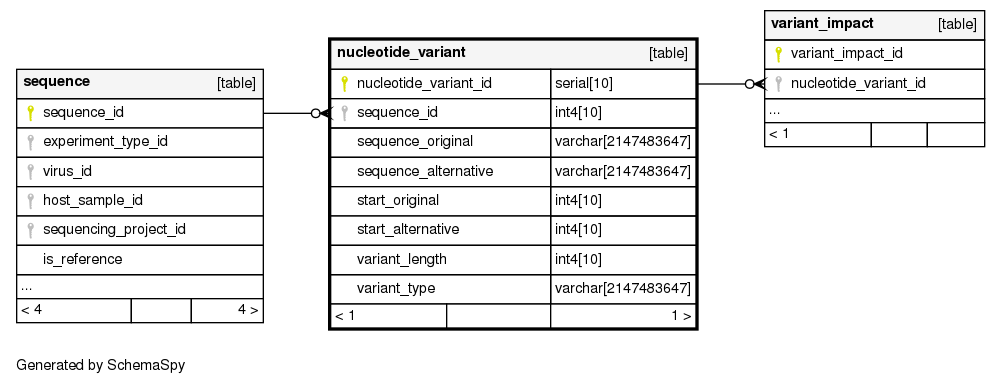
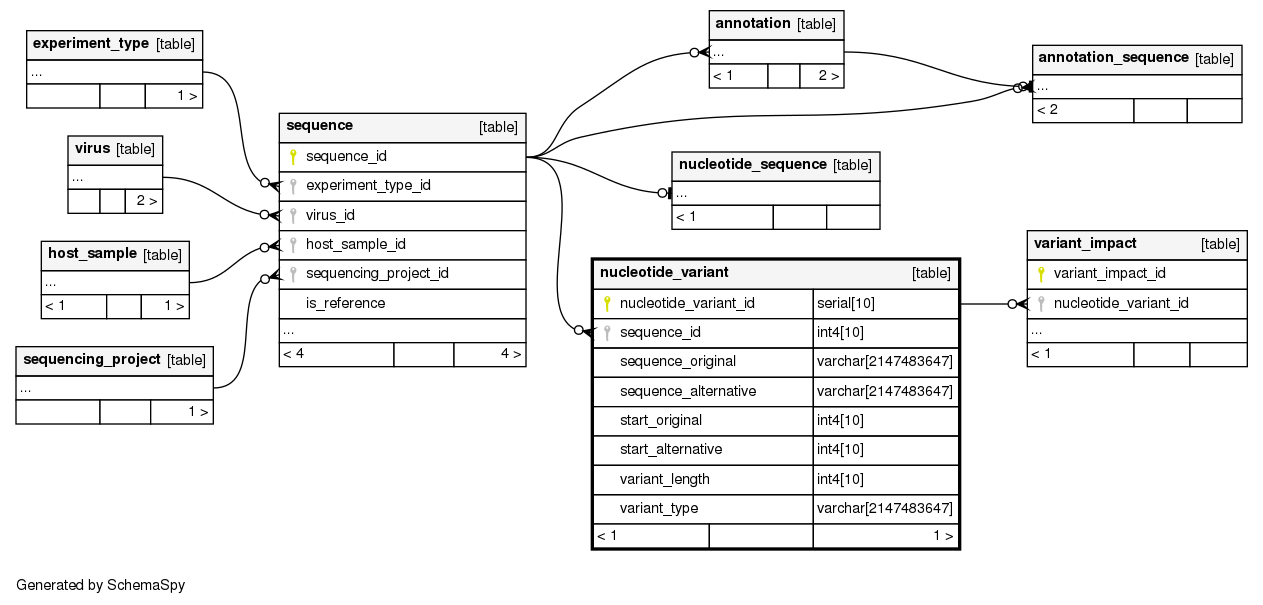
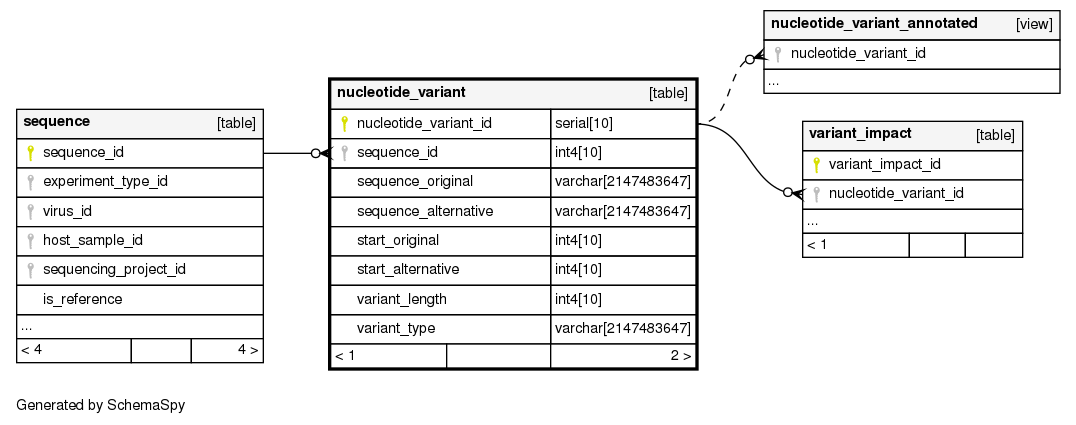
Columns
| Column | Type | Size | Nulls | Auto | Default | Children | Parents | Comments | ||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| nucleotide_variant_id | serial | 10 | √ | nextval('nucleotide_variant_nucleotide_variant_id_seq'::regclass) |
|
|
||||||
| sequence_id | int4 | 10 | null |
|
|
|||||||
| sequence_original | varchar | 2147483647 | null |
|
|
Affected nucleotide sequence from the corresponding reference sequence of the chosen Virus |
||||||
| sequence_alternative | varchar | 2147483647 | null |
|
|
Changed ucleotide sequence (in the target sequence) with respect to the reference one |
||||||
| start_original | int4 | 10 | √ | null |
|
|
Coordinate where the mutation starts on the reference sequrence |
|||||
| start_alternative | int4 | 10 | √ | null |
|
|
Coordinate where the mutation starts on the target sequrence |
|||||
| variant_length | int4 | 10 | null |
|
|
Length of mutation in units of impacted nucleotides |
||||||
| variant_type | varchar | 2147483647 | null |
|
|
Type of nucleotide mutation (SUB = substitution, INS = insertion, DEL = deletion) |
Indexes
| Constraint Name | Type | Sort | Column(s) |
|---|---|---|---|
| nucleotide_variant_pkey | Primary key | Asc | nucleotide_variant_id |
| nuc_var__length | Performance | Asc | variant_length |
| nuc_var__seq_id | Performance | Asc | sequence_id |
| nuc_var__start_alt | Performance | Asc | start_alternative |
| nuc_var__start_orig | Performance | Asc | start_original |